ModernBERT Science DAPT – Experiment Summary
⚠️ Update
A newer version of this model is available:
https://huggingface.co/brettgenz/modernbert-science-dapt-v2
The v2 model was selected using checkpoint evaluation and shows improved retrieval performance and slightly improved classification metrics.
Objective
Evaluate whether domain-adaptive pretraining (DAPT) on a large, curated scientific corpus improves downstream performance of ModernBERT-base on scientific text tasks.
Dataset
Sources
- PMC Open Access (commercial subset)
- arXiv metadata (title + abstract)
Filtering & Policy
- Years: 2015–2024
- Language: English
- License: permissive (CC-BY variants, CC0)
Deduplication
- Exact match removal via SHA256 hash of normalized text
- Conflict resolution:
- Prefer longer documents (char_len)
- Prefer PMC over arXiv
- Stable ordering fallback
Final Training Corpus
- Total documents: 518,531
- arXiv: 276,126
- PMC: 242,405
- Token length (sample):
- median: ~275 tokens
- p95: ~506 tokens
Training Setup
- Base model: answerdotai/ModernBERT-base
- Context length: up to 8192 (padding disabled)
- Framework: Hugging Face Trainer
- Device: Apple Silicon (MPS)
Final Training Run (Run 2)
- Steps: 20,000
- Train loss: 0.996
- Throughput: ~0.6 steps/sec

The training process was stable, with decreasing loss and well-behaved gradient norms.
Evaluation
Evaluation Strategy
We used three proxy evaluations:
- Held-out MLM loss (intrinsic)
- Title → Abstract retrieval (semantic similarity)
- arXiv category classification (top-15) (supervised proxy)
Evaluation Highlights
Retrieval (Title → Abstract)

| Metric | Base | DAPT | Δ |
|---|---|---|---|
| Recall@1 | 0.279 | 0.338 | +21% |
| Recall@5 | 0.565 | 0.668 | +18% |
| MRR | 0.412 | 0.485 | +18% |
Result: Significant improvement in semantic matching and representation quality.
Held-out MLM Loss
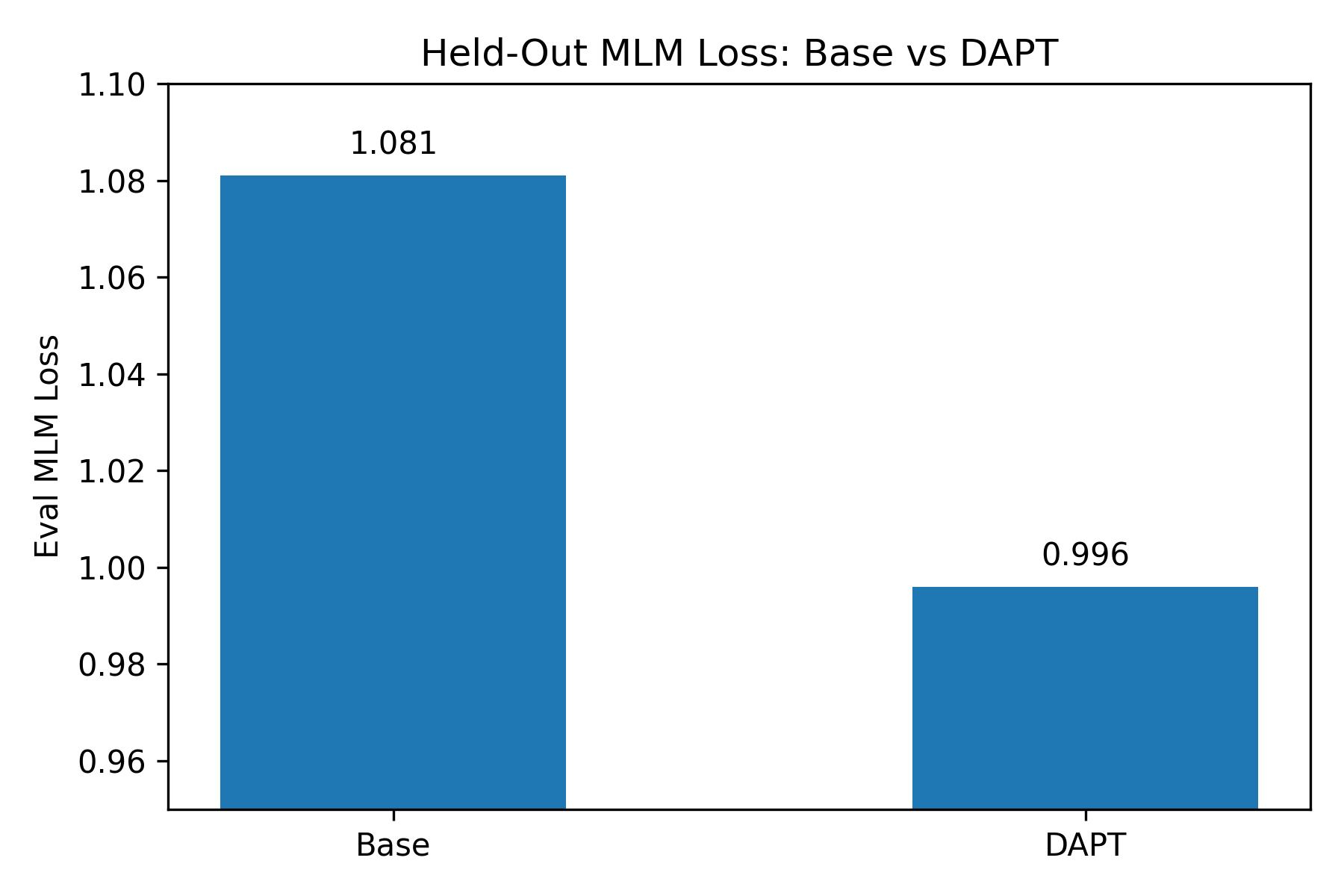
| Model | Eval Loss |
|---|---|
| Base | 1.081 |
| DAPT | 0.996 |
Result: Clear improvement in language modeling on scientific text.
Classification (arXiv Top-15)
| Metric | Base | DAPT |
|---|---|---|
| Accuracy | 0.9161 | 0.9150 |
| Macro F1 | 0.9168 | 0.9156 |
Result: No meaningful change (slight, negligible decrease).
Interpretation
- DAPT successfully improved scientific language understanding, as shown by:
- lower MLM loss
- strong gains in retrieval
- Classification performance remained effectively unchanged, likely due to:
- already high baseline performance (~0.92 macro F1)
- task-specific label boundaries not aligning with DAPT improvements
- retrieval vs classification capturing different capabilities
Conclusion
The DAPT model demonstrates meaningful improvement in scientific-domain representation, particularly for:
- semantic similarity
- retrieval-style tasks
- general language modeling
While classification gains were not observed on this proxy task, there is no evidence of degradation, and improvements in representation quality suggest likely benefit for downstream tasks involving:
- relevance classification
- semantic search
- RAG pipelines
Known Limitations
- PMC data currently skewed toward 2015–2016
- arXiv used only title + abstract (no full text)
- Classification proxy limited to first-category labeling
- No multi-task or instruction tuning applied
Usage
from transformers import AutoTokenizer, AutoModel
model_id = "brettgenz/modernbert-science-dapt-v1"
tokenizer = AutoTokenizer.from_pretrained(model_id)
model = AutoModel.from_pretrained(model_id)
text = "Deep learning approaches for protein structure prediction"
inputs = tokenizer(text, return_tensors="pt", truncation=True)
outputs = model(**inputs)
embeddings = outputs.last_hidden_state
Notes
This model is intended as a drop-in replacement for ModernBERT-base when working with scientific or technical text.
License
This model is released under the Apache 2.0 License.
The training data consists of publicly available open-access scientific content under permissive licenses (e.g., CC-BY, CC0).
- Downloads last month
- 16
Model tree for brettgenz/modernbert-science-dapt-v1
Base model
answerdotai/ModernBERT-base